Are you over 18 and want to see adult content?
More Annotations
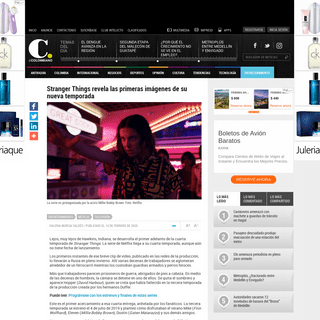
A complete backup of www.elcolombiano.com/entretenimiento/stranger-things-llega-a-su-cuarta-temporada-esto-revelo-en-sus-primera
Are you over 18 and want to see adult content?
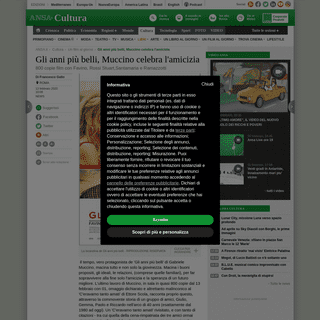
A complete backup of www.ansa.it/sito/notizie/cultura/unfilmalgiorno/2020/02/13/gli-anni-piu-belli-muccino-celebra-lamicizia_d51
Are you over 18 and want to see adult content?

A complete backup of www.hindustantimes.com/education/rsmssb-je-recruitment-2020-1054-vacancies-for-engineers-notified/story-Okd
Are you over 18 and want to see adult content?

A complete backup of hirado.hu/kultura-eletmod/bulvar/cikk/2020/02/14/elkepeszto-sikere-van-az-elso-fotonak-igy-nezni-ki-robert-
Are you over 18 and want to see adult content?
Favourite Annotations

A complete backup of https://touchinglives.org
Are you over 18 and want to see adult content?
A complete backup of https://thepeachkitchen.com
Are you over 18 and want to see adult content?

A complete backup of https://ihergo.com
Are you over 18 and want to see adult content?

A complete backup of https://closeoutexplosion.com
Are you over 18 and want to see adult content?

A complete backup of https://cinemaitaliano.info
Are you over 18 and want to see adult content?
A complete backup of https://golfnow.co.uk
Are you over 18 and want to see adult content?

A complete backup of https://sushisushi-resto.com
Are you over 18 and want to see adult content?
A complete backup of https://freemius.com
Are you over 18 and want to see adult content?
A complete backup of https://mathantics.com
Are you over 18 and want to see adult content?

A complete backup of https://photo-studio9.com
Are you over 18 and want to see adult content?

A complete backup of https://retrohostel.pl
Are you over 18 and want to see adult content?
Text
NEXTSTRAIN
Real-time tracking of pathogen evolution. All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups fromall over the world.
SARS-COV-2 / COVID-19 Further Reading:¶ New China virus: Five questions scientists are asking Nature news 2020-01-22. China virus latest: first US case confirmed Nature news 2020-01-21. New virus surging in Asia rattles scientists Nature news 2020-01-20. New virus identified as likely cause of mystery illness in China Nature news 2020-01-08. Interactive risk analysis by MOBS-lab 2010-01-29INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice. QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.NEXTCLADE
0.14.3 (2021-05-20). The default SARS-CoV-2 reference tree is updated. It allows Nextclade to detect the new Nextstrain clade 21A. See also: Current Nextstrain SARS-CoV-2 clade definitions in nextstrain/ncov GitHub repository: clades.tsv Clade 21A in Nextstrain global build: link Variant 21A/S:154K on CoVariants.org: link Variant 21A/S:478K on CoVariants.org: link RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores at AUSPICE - NEXTSTRAIN Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus thedate range
LOCAL INSTALLATION
Local Installation¶. The following instructions describe how to install augur (bioinformatics tooling) and auspice (our visualization app) together, in the same Conda environment, on macOS or an Ubuntu-style Linux distribution.NEXTCLADE
Clade assignment, mutation calling, and sequence quality checks NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community.NEXTSTRAIN
Real-time tracking of pathogen evolution. All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups fromall over the world.
SARS-COV-2 / COVID-19 Further Reading:¶ New China virus: Five questions scientists are asking Nature news 2020-01-22. China virus latest: first US case confirmed Nature news 2020-01-21. New virus surging in Asia rattles scientists Nature news 2020-01-20. New virus identified as likely cause of mystery illness in China Nature news 2020-01-08. Interactive risk analysis by MOBS-lab 2010-01-29INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice. QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.NEXTCLADE
0.14.3 (2021-05-20). The default SARS-CoV-2 reference tree is updated. It allows Nextclade to detect the new Nextstrain clade 21A. See also: Current Nextstrain SARS-CoV-2 clade definitions in nextstrain/ncov GitHub repository: clades.tsv Clade 21A in Nextstrain global build: link Variant 21A/S:154K on CoVariants.org: link Variant 21A/S:478K on CoVariants.org: link RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores at AUSPICE - NEXTSTRAIN Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus thedate range
LOCAL INSTALLATION
Local Installation¶. The following instructions describe how to install augur (bioinformatics tooling) and auspice (our visualization app) together, in the same Conda environment, on macOS or an Ubuntu-style Linux distribution.NEXTCLADE
Clade assignment, mutation calling, and sequence quality checks QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org. NEXTSTRAIN: ANALYSIS AND VISUALIZATION OF PATHOGEN What is Nextstrain?¶ nextstrain.org aims to provide a real-time snapshot of evolving pathogen populations and to provide interactive data visualizations to virologists, epidemiologists, public health officials, and community scientists. Through interactive data visualizations, we aim to allow exploration of continually up-to-date datasets, providing a novel surveillance tool to the scientific AUSPICE - NEXTSTRAIN.ORG auspice - nextstrain.orgPREPARING YOUR DATA
General formatting tips:¶ The order of the fields doesn’t matter; but if you are going to join your metadata with the global collection then it’s easiest to keep them in the same order!. Not all fields are currently used, but this may change in the future.. Data is case sensitive. The “geographic” columns, such as “region” and “country” will be used to plot the samples on the map. CLADE NAMING & DEFINITIONS Naming¶. We name major clades by the year they are estimated to have emerged and a letter, e.g. 19A, 19B, 20A. The yearly reset of letters will ensure that we don’t progress too far into the alphabet, while the year-prefix provides immediate context on the origin ofAUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
HOW TO INTERPRET THE PHYLOGENETIC TREES Transmission trees vs phylogenetic trees¶. Pathogens spread through rapid replication in one host followed by transmission to another host. An epidemic can only take off when one infection results in more than one subsequent infections. AUSPICE - NEXTSTRAIN Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus thedate range
AUSPICE: AN OPEN-SOURCE INTERACTIVE TOOL FOR VISUALISING This work is made possible by the open sharing of genetic data by research groups from all over the world. We gratefully acknowledge their contributions.AUGUR ANCESTRAL
Named Arguments¶--tree, -t. prebuilt Newick--alignment, -a. alignment in fasta or VCF format--output-node-data. name of JSON file to save mutations and ancestral sequences to NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores atNEXTCLADE
At the core of Nextclade is a pairwise sequence alignment between the reference sequence and each of the sequences in your data. Once the sequences are aligned, we can identify differences between them and use these differences to place the sequence on the AUSPICE - NEXTSTRAIN auspice. Date Range. Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus the date range here can be wider than the sample collection range. 2019-12-19. 2021-05-30. Color By. Change theAUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
EXPLORE ZIKA VIRUS EVOLUTION Explore Zika virus evolution¶. This tutorial explains how to create a Nextstrain build for the Zika virus. We will first make the build step-by-step using an example data set. Then we will see how to automate this stepwise process by defining a pathogen build script. AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores atNEXTCLADE
At the core of Nextclade is a pairwise sequence alignment between the reference sequence and each of the sequences in your data. Once the sequences are aligned, we can identify differences between them and use these differences to place the sequence on the AUSPICE - NEXTSTRAIN auspice. Date Range. Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus the date range here can be wider than the sample collection range. 2019-12-19. 2021-05-30. Color By. Change theAUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
EXPLORE ZIKA VIRUS EVOLUTION Explore Zika virus evolution¶. This tutorial explains how to create a Nextstrain build for the Zika virus. We will first make the build step-by-step using an example data set. Then we will see how to automate this stepwise process by defining a pathogen build script. AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice. AUSPICE - NEXTSTRAIN.ORG auspice - nextstrain.orgLOCAL INSTALLATION
Local Installation¶. The following instructions describe how to install augur (bioinformatics tooling) and auspice (our visualization app) together, in the same Conda environment, on macOS or an Ubuntu-style Linux distribution. HOW TO INTERPRET THE PHYLOGENETIC TREES Interpreting these should, however, be done with caution, as the sampling and sequencing or lack thereof can significantly influence the interpretation, In the following example, we first show fully sampled phylogenetic tree, with samples from two different locations denoted by orange and blue. In the fully sampled case on the right,our
AUGUR TREE — AUGUR DOCUMENTATION Currently, only available for IQTREE. Default: “GTR”. --nthreads. number of threads to use; specifying the value ‘auto’ will cause the number of available CPU cores on your system, if determinable, to be used. Default: 1. --vcf-reference. fasta file of the sequence the VCF was mapped to. --exclude-sites. file name of one-based sites toAUGUR ALIGN
If gaps represent missing data rather than true indels, replace by N after aligning. An existing alignment to which the sequences will be added. The ouput alignment will be the same length as this existing alignment. Produce extra files (e.g. pre- and post-aligner files) AUSPICE: AN OPEN-SOURCE INTERACTIVE TOOL FOR VISUALISING Auspice is software to display beautiful, interactive visualisations of phylogenomic data. ¶. Auspice being used to show the spread of influenza H7N9 virus across Asia. Communicating scientific results while also allowing interrogation of the underlying data is an integral part of the scientific process. Current scientific publishingpractices
AUGUR FREQUENCIES
date to end frequencies calculations; may be specified as an Augur-style numeric date (with the year as the integer part) or YYYY-MM-DD. --tree, -t. tree to estimate clade frequencies for. --include-internal-nodes. calculate frequencies for internal nodes as well as tips. Default: False. NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores atNEXTCLADE
At the core of Nextclade is a pairwise sequence alignment between the reference sequence and each of the sequences in your data. Once the sequences are aligned, we can identify differences between them and use these differences to place the sequence on the AUSPICE - NEXTSTRAIN auspice. Date Range. Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus the date range here can be wider than the sample collection range. 2019-12-19. 2021-05-30. Color By. Change theAUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
EXPLORE ZIKA VIRUS EVOLUTION Explore Zika virus evolution¶. This tutorial explains how to create a Nextstrain build for the Zika virus. We will first make the build step-by-step using an example data set. Then we will see how to automate this stepwise process by defining a pathogen build script. AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores atNEXTCLADE
At the core of Nextclade is a pairwise sequence alignment between the reference sequence and each of the sequences in your data. Once the sequences are aligned, we can identify differences between them and use these differences to place the sequence on the AUSPICE - NEXTSTRAIN auspice. Date Range. Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus the date range here can be wider than the sample collection range. 2019-12-19. 2021-05-30. Color By. Change theAUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
EXPLORE ZIKA VIRUS EVOLUTION Explore Zika virus evolution¶. This tutorial explains how to create a Nextstrain build for the Zika virus. We will first make the build step-by-step using an example data set. Then we will see how to automate this stepwise process by defining a pathogen build script. AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice. AUSPICE - NEXTSTRAIN.ORG auspice - nextstrain.orgLOCAL INSTALLATION
Local Installation¶. The following instructions describe how to install augur (bioinformatics tooling) and auspice (our visualization app) together, in the same Conda environment, on macOS or an Ubuntu-style Linux distribution. HOW TO INTERPRET THE PHYLOGENETIC TREES Interpreting these should, however, be done with caution, as the sampling and sequencing or lack thereof can significantly influence the interpretation, In the following example, we first show fully sampled phylogenetic tree, with samples from two different locations denoted by orange and blue. In the fully sampled case on the right,our
AUGUR TREE — AUGUR DOCUMENTATION Currently, only available for IQTREE. Default: “GTR”. --nthreads. number of threads to use; specifying the value ‘auto’ will cause the number of available CPU cores on your system, if determinable, to be used. Default: 1. --vcf-reference. fasta file of the sequence the VCF was mapped to. --exclude-sites. file name of one-based sites toAUGUR ALIGN
If gaps represent missing data rather than true indels, replace by N after aligning. An existing alignment to which the sequences will be added. The ouput alignment will be the same length as this existing alignment. Produce extra files (e.g. pre- and post-aligner files) AUSPICE: AN OPEN-SOURCE INTERACTIVE TOOL FOR VISUALISING Auspice is software to display beautiful, interactive visualisations of phylogenomic data. ¶. Auspice being used to show the spread of influenza H7N9 virus across Asia. Communicating scientific results while also allowing interrogation of the underlying data is an integral part of the scientific process. Current scientific publishingpractices
AUGUR FREQUENCIES
date to end frequencies calculations; may be specified as an Augur-style numeric date (with the year as the integer part) or YYYY-MM-DD. --tree, -t. tree to estimate clade frequencies for. --include-internal-nodes. calculate frequencies for internal nodes as well as tips. Default: False. NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice. RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores atNEXTCLADE
At the core of Nextclade is a pairwise sequence alignment between the reference sequence and each of the sequences in your data. Once the sequences are aligned, we can identify differences between them and use these differences to place the sequence on theINSTALLATION
Docker¶. Docker is a very popular container system freely-available for all platforms. When you use Docker with the Nextstrain CLI, you don’t need to install any other Nextstrain software dependencies as validated versions are already bundled into a container image by theNextstrain team.
HOW TO INTERPRET THE PHYLOGENETIC TREES Interpreting these should, however, be done with caution, as the sampling and sequencing or lack thereof can significantly influence the interpretation, In the following example, we first show fully sampled phylogenetic tree, with samples from two different locations denoted by orange and blue. In the fully sampled case on the right,our
CUSTOMIZING YOUR AUSPICE VISUALIZATION Customizing your Auspice visualization. Just as we can specify a build-specific analysis options in the builds.yaml file, we can also specify build-specific visualization options in this directory. Looking at the builds.yaml file, the last few lines are: This points to a JSON file that parameterizes the output files used forvisualizion with
NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice. RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores atNEXTCLADE
At the core of Nextclade is a pairwise sequence alignment between the reference sequence and each of the sequences in your data. Once the sequences are aligned, we can identify differences between them and use these differences to place the sequence on theINSTALLATION
Docker¶. Docker is a very popular container system freely-available for all platforms. When you use Docker with the Nextstrain CLI, you don’t need to install any other Nextstrain software dependencies as validated versions are already bundled into a container image by theNextstrain team.
HOW TO INTERPRET THE PHYLOGENETIC TREES Interpreting these should, however, be done with caution, as the sampling and sequencing or lack thereof can significantly influence the interpretation, In the following example, we first show fully sampled phylogenetic tree, with samples from two different locations denoted by orange and blue. In the fully sampled case on the right,our
CUSTOMIZING YOUR AUSPICE VISUALIZATION Customizing your Auspice visualization. Just as we can specify a build-specific analysis options in the builds.yaml file, we can also specify build-specific visualization options in this directory. Looking at the builds.yaml file, the last few lines are: This points to a JSON file that parameterizes the output files used forvisualizion with
NEXTSTRAIN: ANALYSIS AND VISUALIZATION OF PATHOGEN Nextstrain is a collection of open-source tools to aid in our understanding of pathogen spread and evolution, especially in outbreak scenarios. We have designed these in such a way that they can be used with a wide range of data sources, and are easy to replace with your own tooling. Broadly speaking, Nextstrain consists of. LEARN — NEXTSTRAIN DOCUMENTATION Table of contents. About Nextstrain. Interpreting Nextstrain. How to interpret the phylogenetic trees. Interacting with Nextstrain’s phylogeographic visualisations. Interpreting the Map. Pathogens.PREPARING YOUR DATA
General formatting tips:¶ The order of the fields doesn’t matter; but if you are going to join your metadata with the global collection then it’s easiest to keep them in the same order!. Not all fields are currently used, but this may change in the future.. Data is case sensitive. The “geographic” columns, such as “region” and “country” will be used to plot the samples on the map. AUSPICE - NEXTSTRAIN.ORG auspice - nextstrain.org CLADE NAMING & DEFINITIONS Clade Naming & Definitions. The nomenclature used by Nextstrain to designate clades for SARS-CoV-2 is driven by the following objectives: label genetically well defined clades that have reached significant frequency and geographic spread, allow for transient clade designations that are elevated to major clades if they persist andrise in
USING AUGUR
Using Augur ¶. Using Augur. Augur consists of a number of tools that allow the user to filter and align sequences, infer trees, and integrate the phylogenetic analysis with meta data. The different tools are meant to be composable and the output of one command will serve as the input of other commands. All of Augur’s commands areaccessed
WRITING A NARRATIVE
Step 2: Start a Simple Narrative ¶. We’re going to start by creating a narrative with only one page (the title page). mkdir narratives touch narratives/example.md. Open up narratives/example.md and start by pasting in the following YAML frontmatter, which is used to define some basic set-up for the narrative and create the firstnarrative page.
AUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org. RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores at AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
AUGUR ALIGN
If gaps represent missing data rather than true indels, replace by N after aligning. An existing alignment to which the sequences will be added. The ouput alignment will be the same length as this existing alignment. Produce extra files (e.g. pre- and post-aligner files)AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
AUGUR FREQUENCIES
date to end frequencies calculations; may be specified as an Augur-style numeric date (with the year as the integer part) or YYYY-MM-DD. --tree, -t. tree to estimate clade frequencies for. --include-internal-nodes. calculate frequencies for internal nodes as well as tips. Default: False. NEXTSTRAINAUSPICEDOCSTUBERCULOSISAVIAN INFLUENZAMUMPSWEST NILE VIRUS Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response.NEXTSTRAIN
All source code is freely available under the terms of the GNU Affero General Public License.Screenshots may be used under a CC-BY-4.0 license and attribution to nextstrain.org must be provided.. This work is made possible by the open sharing of genetic data by research groups from all over the world. SARS-COV-2 / COVID-19 The virus has been named SARS-CoV-2 and the disease it causes is COVID-19. As of March 6, 2020 over 100,685 cases and 3,411 deaths have been reported . The WHO currently estimates the case fatality rate at 3.4%, which is significantly less than the case fatality rate of SARS. The case counts are dramatically rising in part due to increased QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org. RUNNING THE ANALYSIS --profile tells snakemake where to find your builds.yaml and config.yaml files.-p tells snakemake to print each command it runs to help you understand what it’s doing.. If you’d like to run a dryrun, try running with the -np flag, which will execute a dryrun. This prints out each command, but doesn’t execute it. Note that the example profile runs the workflow with at most two cores at AUGUR RELEASES & UPGRADING Augur Releases & Upgrading¶. To make an omelette, you have to break some eggs. To keep improving and adding to augur and auspice, we sometimes have to break old code!. We know these kinds of changes can be really frustrating for users - things that used to work suddenlydon’t.
AUGUR REFINE
Specify the evolutionary rate¶. By default augur (through treetime) will estimate the rate of evolution from the data by regressing divergence vs sampling date.In some scenarios, however, there is insufficient temporal signal to reliably estimate the rate and the analysis will be more robust and reproducible if one fixes this rateexplicitly.
AUGUR ALIGN
If gaps represent missing data rather than true indels, replace by N after aligning. An existing alignment to which the sequences will be added. The ouput alignment will be the same length as this existing alignment. Produce extra files (e.g. pre- and post-aligner files)AUGUR ANCESTRAL
Possible choices: joint, marginal. calculate joint or marginal maximum likelihood ancestral sequence states. Default: “joint”. --vcf-reference. fasta file of the sequence the VCF was mapped to. --output-vcf. name of output VCF file which will include ancestral seqs. --keep-ambiguous. do not infer nucleotides at ambiguous (N)sites on tip
AUGUR FREQUENCIES
date to end frequencies calculations; may be specified as an Augur-style numeric date (with the year as the integer part) or YYYY-MM-DD. --tree, -t. tree to estimate clade frequencies for. --include-internal-nodes. calculate frequencies for internal nodes as well as tips. Default: False. QUICKSTART — NEXTSTRAIN DOCUMENTATION Quickstart¶. This guide uses the Nextstrain command-line interface (CLI) to help you get started running and viewing the pathogen builds you see on nextstrain.org.It assumes you are comfortable using the command line and installing software on your computer. If you need help when following this guide, please reach out by emailing us, or create a post at discussion.nextstrain.org.INSTALL NEXTSTRAIN
Note. If you want to contribute to the development of Nextstrain tools, see the developer documentation for instructions to install these tools from source.. If you prefer to manage your own custom environment (e.g., a Conda environment, Docker image, environment modules on a cluster, etc.), see the individual installation documentation for Nextstrain CLI, Augur, or Auspice.PREPARING YOUR DATA
General formatting tips:¶ The order of the fields doesn’t matter; but if you are going to join your metadata with the global collection then it’s easiest to keep them in the same order!. Not all fields are currently used, but this may change in the future.. Data is case sensitive. The “geographic” columns, such as “region” and “country” will be used to plot the samples on the map. AUSPICE - NEXTSTRAIN auspice. Phylogeny of SARS-like betacoronaviruses including novel coronavirus SARS-CoV-2. Built with blab/sars-like-cov. Maintained byTrevor Bedford and
AUSPICE - NEXTSTRAIN auspice. Date Range. Use this slider to filter the data based on the sample date. This may include inferred dates for ancestral nodes in the tree, and thus the date range here can be wider than the sample collection range. 2019-12-19. 2021-05-30. Color By. Change theUSING AUGUR
Using Augur ¶. Using Augur. Augur consists of a number of tools that allow the user to filter and align sequences, infer trees, and integrate the phylogenetic analysis with meta data. The different tools are meant to be composable and the output of one command will serve as the input of other commands. All of Augur’s commands areaccessed
AUGUR FILTER
As an example, we’ll look that the filter command in greater detail using material from the Zika tutorial . The filter command allows you to selected various subsets of your input data for different types of analysis. A simple example use of this command would be. augur filter --sequences data/sequences.fasta --metadata data/metadata.tsvAUGUR ALIGN
If gaps represent missing data rather than true indels, replace by N after aligning. An existing alignment to which the sequences will be added. The ouput alignment will be the same length as this existing alignment. Produce extra files (e.g. pre- and post-aligner files) HOW TO INTERPRET THE PHYLOGENETIC TREES Interpreting these should, however, be done with caution, as the sampling and sequencing or lack thereof can significantly influence the interpretation, In the following example, we first show fully sampled phylogenetic tree, with samples from two different locations denoted by orange and blue. In the fully sampled case on the right,our
AUSPICE: AN OPEN-SOURCE INTERACTIVE TOOL FOR VISUALISING Auspice is software to display beautiful, interactive visualisations of phylogenomic data. ¶. Auspice being used to show the spread of influenza H7N9 virus across Asia. Communicating scientific results while also allowing interrogation of the underlying data is an integral part of the scientific process. Current scientific publishingpractices
Nextstrain
HELP
DOCS
BLOG LOGIN
Nextstrain
Real-time tracking of pathogen evolution Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community. Our goal is to aid epidemiological understanding and improve outbreak response. If you have any questions, or simply want to say hi, please give us a shout at helloobfuscate@nextstrain.org.Read More
SARS-CoV-2 (COVID-19) We are incorporating SARS-CoV-2 genomes as soon as they are shared and providing analyses and situation reports. In addition we have developed a number of resources and tools, and are facilitating independent groups to run their own analyses. Please see the SARS-CoV-2 resources page for more information. All datasets and resources Latest global analysis Latest Situation Report 2020-08-14Explore pathogens
Genomic analyses of specific pathogens kept up-to-date by theNextstrain team
SARS-CoV-2
Seasonal influenza
West Nile virus
Mumps
Zika
West African Ebola 2013-16Dengue
Avian influenza
Measles
Enterovirus D68
Tuberculosis
From the community
Analyses by independent groups stored and accessed via public GitHubrepos
DRC Ebola (2018-19)
Chikungunya
Lassa in Nigeria (2015-18)RSV (subtype A)
Cassava geminivirusesWheat yellow rust
Narratives
Narratives are a method of data-driven storytelling. They allow authoring of content which is displayed alongside a view into the data. See here to find out more. nCoV Situation Report 2020-05-15 20 years of WNV in the Americas Example Situation Report for current DRC Ebola outbreakPhilosophy
Pathogen Phylogenies In the course of an infection and over an epidemic, pathogens naturally accumulate random mutations to their genomes. This is an inevitable consequence of error-prone genome replication. Since different genomes typically pick up different mutations, mutations can be used as a marker of transmission in which closely related genomes indicate closely related infections. By reconstructing a _phylogeny_ we can learn about important epidemiological phenomena such as spatial spread, introduction timings and epidemic growth rate. Actionable Inferences However, if pathogen genome sequences are going to inform public health interventions, then analyses have to be rapidly conducted and results widely disseminated. Current scientific publishing practices hinder the rapid dissemination of epidemiologically relevant results. We thought an open online system that implements robust bioinformatic pipelines to synthesize data from across research groups has the best capacity to make epidemiologically actionable inferences.This Website
This website aims to provide a _real-time_ snapshot of evolving pathogen populations and to provide interactive data visualizations to virologists, epidemiologists, public health officials and citizen scientists. Through interactive data visualizations, we aim to allow exploration of continually up-to-date datasets, providing a novel surveillance tool to the scientific and public health communities.Future Directions
Nextstrain is under active development and we have big plans for its future, including visualization, bioinformatics analysis and an increasing number and variety of datasets. If you have any questions or ideas, please give us a shout at helloobfuscate@nextstrain.org. A bioinformatics and data viz toolkit Nextstrain provides an open-source toolkit enabling the bioinformatics and visualization you see on this site. Tweak our analyses and create your own using the same tools we do. We aim to empower the wider genomic epidemiology and public health communities. Read the documentation Hadfield _et al., _Nextstrain: real-time tracking of pathogenevolution _,
Bioinformatics_ (2018) Nextstrain is built byTrevor Bedford
,Richard Neher
,James
Hadfield ,Emma
Hodcroft ,Thomas Sibley,John
Huddleston ,Ivan Aksamentov,Jover Lee
,Kairsten Fay
,Moira Zuber,Eli Harkins ,Misja Ilcisin ,Cassia Wagner ,Louise Moncla ,Allison Black ,Sidney Bell ,Miguel Parades,Colin Megill
,Barney Potter
,Pavel
Sagulenko ,Charlton Callender All source code is freely available under the terms of the GNU Affero General Public License.
Screenshots may be used under a CC-BY-4.0 licenseand attribution to
nextstrain.org must be provided. This work is made possible by the open sharing of genetic data by research groups from all over the world. We gratefully acknowledge their contributions. Special thanks to Kristian Andersen, Josh Batson, David Blazes, Jesse Bloom, Peter Bogner, Anderson Brito, Matt Cotten, Ana Crisan, Tulio de Oliveira, Gytis Dudas, Vivien Dugan, Karl Erlandson, Nuno Faria, Jennifer Gardy, Nate Grubaugh, Becky Kondor, Dylan George, Ian Goodfellow, Betz Halloran, Christian Happi, Jeff Joy, Paul Kellam, Philippe Lemey, Nick Loman, Duncan MacCannell, Erick Matsen, Sebastian Maurer-Stroh, Placide Mbala, Danny Park, Oliver Pybus, Andrew Rambaut, Colin Russell, Pardis Sabeti, Katherine Siddle, Kristof Theys, Dave Wentworth, Shirlee Wohl and Cecile Viboud for comments, suggestions and data sharing. Splash page images stylised in Lunapic . Zikadrawing
by David Goodwill, Dengue EM by Zhang _et al., _Ebola EM by Frederick Murphy / CDC, Seasonal Influenza,
Lassa and
West Nile Virus images by Cynthia Goldsmith / CDC, Avian Influenza (A/H7N9) by Cynthia Goldsmith and Thomas Rowe / CDC, Mumpsby the CDC, Measles
by Shmuel Rozenblatt, Enterovirus by Shingler _et al., _Tuberculosisby Ray Butle, RSV
by NIAID / NIH, Cassava Leaf by Wilmer Cuellar, Wheat yellow rust (Puccinia striiformis)by
Yue Jin / USDA, coronavirusby
Dr. Fred Murphy / CDC. Nextstrain is supported by � 2015-2021 Trevor Bedford and Richard NeherDetails
Copyright © 2024 ArchiveBay.com. All rights reserved. Terms of Use | Privacy Policy | DMCA | 2021 | Feedback | Advertising | RSS 2.0